Fabrizio Fasano1,2, John Evans3, Chloe Benson4, Yifei Wang4, Derek K Jones3,5, Alison Paul4, and Robert Turner6,7
1Siemens Healthcare Ltd, Camberly, United Kingdom, 2Siemens Healthcare GmbH, Erlangen, Germany, 3Cardiff University Brain Research Imaging Centre (CUBRIC), Cardiff University, Cardiff, United Kingdom, 4School of Chemistry, Cardiff University, Cardiff, United Kingdom, 5Mary McKillop Institute for Health Research, Faculty of Health Sciences, Australian Catholic University, Melbourne, Australia, 6Department of Neurophysics, Max Planck Institute for Human Cognitive and Brain Sciences, Leipzig, Germany, 7Sir Peter Mansfield Imaging Centre, University of Nottingham, Nottingham, United Kingdom
1Siemens Healthcare Ltd, Camberly, United Kingdom, 2Siemens Healthcare GmbH, Erlangen, Germany, 3Cardiff University Brain Research Imaging Centre (CUBRIC), Cardiff University, Cardiff, United Kingdom, 4School of Chemistry, Cardiff University, Cardiff, United Kingdom, 5Mary McKillop Institute for Health Research, Faculty of Health Sciences, Australian Catholic University, Melbourne, Australia, 6Department of Neurophysics, Max Planck Institute for Human Cognitive and Brain Sciences, Leipzig, Germany, 7Sir Peter Mansfield Imaging Centre, University of Nottingham, Nottingham, United Kingdom
The multi-shot inversion recovery slice shuffled EPI approach, recently proposed by Sanchez and co-workers to map myelin patterns, shows good reproducibility and good quality point spread function, appearing suitable to detect myelination processes at a nominal 400μm in-plane resolution.
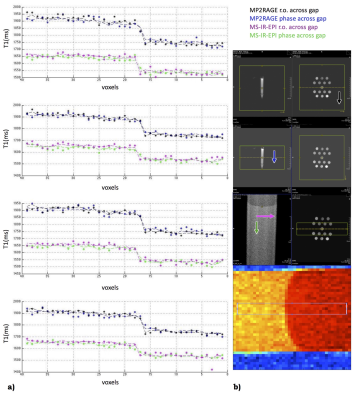
Fig.3. Signal profiles of 4 lines across the gel boundary. Dashed lines represent the average values across the lines. Voxels size is 400µm; b) From top: legend, positioning for MP2RAGE and MS-IR-EPI, the 4 lines sampling. Arrows indicate phase (phase 2 for MP2RAGE) direction. Both MPRAGE and MS-IR-EPI show a good profile, mostly affected by the true object shape/partial volume/k-space sampling effect. The difference in T1 estimation is expected, being MP2RAGE protocol suboptimal to assess a 1500ms T1 (our focus is on PSF here).

Fig.4. OTF, PSF and signal intensity profile estimated for an image profile with a gel boundary thickness similar to that of phantom #14. T1=1500ms. The true profile is superimposed. For sake of simplicity, full-Fourier versions were simulated. The estimated MS-IR-EPI and TI=2250 MP2RAGE profiles are close to the true profile. The estimated PSF and signal profile of MP2RAGE for TI=800ms show significant broadening, incompatible with imaging typical cortical myelinated layers
